 
You can also load features that can include additional sample data, ex. You should read more about this in the CoNet manual so that your lineage and taxon columns are formatted correctly (if using). The metadata here is a table including OTU names that match the infile, along with groups, lineage, and taxon columns. The infile is essentially a modified OTU table with OTUs in rows and samples in columns, note that OTUs from each 'group' should be present as their own row. You can refer to the CoNet tutorial page to see examples of input and metadata files. First, you will need to prepare your infiles. Now that you can run small analyses, try running CoNet with your own files. Feel free to change the layout as described above. After hitting GO, a graph should show up in the Cytoscape window. If you follow the 'long version' instructions, you can start to get a feel for how to navigate through all the CoNet menus. Next follow short version or long version instructions closely. Use wget to grab the Costello tutorial files available from. Now try loading settings from the tutorial files. circular, or organic (this one needs to be installed via the menu options before it can be used). You can change the layout of this graph to your liking by clicking on the 'layout' tab in Cytoscape, and picking an alternative layout, ex. After a moment, you should see a graph show up in the Cytoscape window. Test that CoNet is working with Cytoscape, by clicking 'demo' in the CoNet app. 
Click on the 'Apps' tab in Cytoscape, then click on the CoNet app. Launch CoNet from Cytoscape to use the graphical user interface to set up your initial run settings. Load the CoNet plug in for Cytoscape by clicking on the 'Apps' tab in Cytoscape, install from file, then browse to your CoNet installation /path/to/CoNet3/conet.jar. It will open up a graphical user interface. Test CoNet, start it using the alias set above and use -h to see the help. CoNet is very memory intensive.Įxport CLASSPATH=$/CoNet.jarĪlias conet="java -Xmx800m be.ac.CooccurrenceAnalyser" 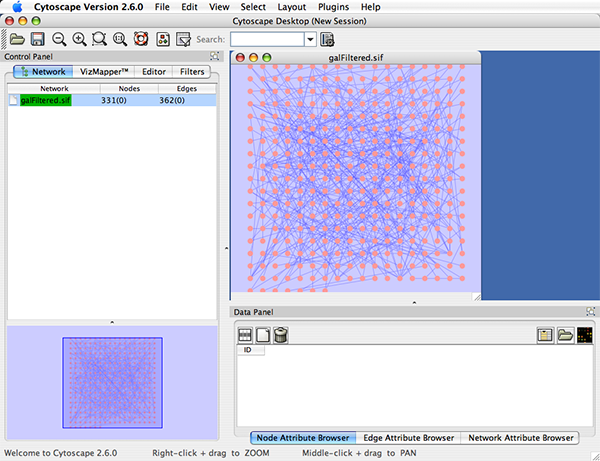 
You can allocate more memory to java by setting the -Xmx to a higher number according to how much memory you have on your system, ex. Set environment variables (and an alias if you like) in your. I'm assuming you already have miniconda3 or anaconda on your system.Ĭonda create -name cytoscape -c bioconda cytoscape Set up a new conda environment called cytoscape. CoNet3 needs Cytoscape 3 that needs at least Java 7 (openjdk 1.7.0+). I'm assuming you're working in linux at the command line. Step-by-step method for using CoNet with CytoscapeĬheck what version of java you have installed. Here are the steps I took to get everything working right. Since I was having trouble customizing the analysis in R I switched to Cystoscape instead for more control. Using CoNet in R was a little challenging as the documentation is very abbreviated. CytoNet is available as a 'barebonesCoNet' function in the seqgroup package in R, but the full functionality is only available through the CoNet plug-in that works with Cytoscape (Faust and Raes, 2016 Faust, 2021). False positives due to compositional effects are controlled through permutation and bootstrapping. CytoNet is an ensemble network analysis method that combines several similarity and dissimilarity methods into a single tool (Faust et al., 2012). 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
